BioSure® Trout Erythrocyte Nuclei (TENs) are an internal biological reference material that can function as a DNA standard when determining nuclear DNA content of human, animal, or plant cells by flow cytometry1 2 3 4. The DNA content of rainbow trout red blood cells (5.149 - 5.240 pg) has been reported to be 80% of human diploid cells (6.436 - 6.550 pg), and consequently may provide a better internal DNA standard than chicken red blood cells1.
TENs are isolated from fresh rainbow trout blood, Oncorhynchus mykiss, and fixed in ethanol-PBS. The procedure yields single nuclei, doublets, triplets, and a few larger aggregates. DNA linear scale histograms of propidium iodide (PI) stained nuclei exhibit multiple peaks, single nuclei constitute peak one. Other performance characteristics for this product have not been established.
FORM
BioSure® TENs are supplied as an unstained suspension in 2.0 milliliter volumes. The nuclei count per milliliter is at least 2.0 X 107.
STABILITY
TENs are stable at 2° - 8° C until the expiration date denoted on each vial label.
PREPARATION OF PROPIDIUM IODIDE (50 microgram/mL) STAINING SOLUTION
To 48 milliliters of calcium and magnesium free Dulbecco's PBS, add 2.5 mg of propidium iodide (PI) and 0.3 mL of Nonidet P40 (NP40). Mix the solution and Q.S. to 50 mL. Staining solution is stable at 2° - 8° C for at least 30 days when stored protected from light in an amber bottle.
PREPARATION OF WORKING SOLUTION
The recommended working solution is a 1:10 dilution of the stock TENs in the PI stain prepared previously. This provides an adequate density of nuclei and sufficient working solution for several hours.
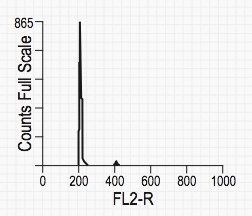
Typical TEN graph
1. Gently mix the TEN stock for 5 - 10 seconds to suspend the nuclei, and add two drops to a test tube (13 X 75 mm).
2. Add 1.0 mL of PI stain, mix for 5 - 10 seconds, and incubate at room temperature (22° - 25° C) for ten minutes protected from light.
3. Mix sample 5 - 10 seconds and evaluate. Samples are typically stable at 2° - 8° C for four hours when protected from light.
NOTE: The flow cell of some cytometers is susceptible to clogging. Filtering the stained preparation through a 30 - 50 mesh nylon screen will eliminate large nuclear aggregates.
LITERATURE CITED
1Vindelov LL, Christensen IJ, Nissen NI: Standardization of High-Resolution Flow Cytometric DNA Analysis by the Simultaneous Use of Chicken and Trout Red Blood Cells as Internal Reference Standards. Cytometry 3: 328, 1983.
2Vindelov LL, Christensen IJ, Jensen G, Nissen NI: Limits of Detection of Nuclear DNA Abnormalities by Flow Cytometric DNA Analysis. Results Obtained by a Set of Methods for Sample-Storage, Staining and Internal Standardization. Cytometry 3: 332, 1983.
3Petersen SE: Accuracy and Reliability of Flow Cytometric DNA Analysis Using a Simple, One-Step Ethidium Bromide Staining Protocol. Cytometry 7: 301, 1986.
4Gregory, T.R.; Animal Genome Size Database: 2001-2005, available at http://www.genomesize.com